In Silico Assessment of Preclinical Disease Models
industrial doctorate, preclinical models, single-cell RNA-seq, singIST, atopic dermatitis, skin disease, Almirall, computational biology, drug development, UPC
| Type | Industrial Doctorate (Doctorat Industrial) |
| Doctoral student | Aitor Moruno-Cuenca |
| Supervisor | Alexandre Perera Lluna |
| Company partner | Almirall SA (formerly Laboratoris Almirall SA) |
| University | Universitat Politècnica de Catalunya (UPC) |
| Reference | futur.upc.edu/37915145 |
| Status | Active |
Context: A long-standing problem in drug development
One of the most persistent frustrations in biomedical research is a simple, costly mismatch: a treatment that works in mice often fails in humans. A major reason is that the animal models used to test drug candidates are selected largely based on qualitative judgement — shared symptoms, broad gene expression overlap, or historical convention — rather than rigorous, quantitative comparison to the human disease.
The consequence is systematic: if the model does not capture the biology that the drug targets, the experiment tells you nothing useful, regardless of how well the drug performs. This is a major driver of high attrition rates in clinical trials, particularly in complex inflammatory and immune-mediated diseases.
Almirall SA is a Barcelona-headquartered pharmaceutical company specialising in dermatology and immunology-mediated skin diseases. Their pipeline includes treatments for psoriasis, atopic dermatitis, hidradenitis suppurativa, and other conditions where preclinical modelling is both essential and notoriously difficult. The need for better, more principled methods to evaluate animal models before committing to costly development programmes is a direct industrial challenge that this doctorate addresses.
Research: singIST — a quantitative framework for model comparison
The central contribution of this industrial doctorate is singIST (single-cell Integrative Similarity Tool), a computational method that quantifies how faithfully a preclinical disease model reproduces the biology of the corresponding human disease — at the resolution of individual cell types, biological pathways, and genes.
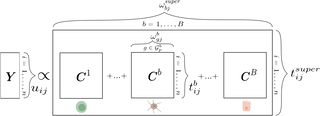
How singIST works
singIST takes single-cell RNA sequencing (scRNA-seq) data from both a preclinical model (typically mouse) and human patients and computes similarity at three nested levels:
The method addresses two biological realities that simpler approaches ignore:
- Gene conservation: not every human gene has a functional mouse equivalent; singIST accounts for this explicitly rather than silently mapping non-conserved genes.
- Cell type composition: the relative abundance of cell populations differs between species; a cell type that is common in human disease may be rare or absent in the mouse model.
The result is a set of interpretable similarity scores that allow statements like: “this model captures keratinocyte biology well but misses the dendritic cell response entirely” — rather than just a broad verdict of “similar” or “different”.
Validation on skin disease models
singIST was validated on three mouse models of atopic dermatitis (AD) — a common inflammatory skin condition — and applied to data from human hidradenitis suppurativa (HS), a chronic inflammatory skin disease. These conditions sit squarely within Almirall’s therapeutic focus.
The method reproduced established knowledge (correctly identifying which mouse model best captures the Th2-skewed immune response characteristic of human AD) while generating novel hypotheses about specific pathways and cell types that each model captures or misses. The discriminative power — the ability to distinguish between models that seem superficially similar — is one of the tool’s key practical strengths.
Why this matters for industrial drug development
The tool is freely available and integrates with standard scRNA-seq pipelines — no additional data collection is required beyond what a well-equipped computational biology team already produces.
Publication
Related blog post: Does your mouse model actually look like a human disease? Now there’s a tool to check, cell by cell
About the partners
Almirall SA is a Barcelona-based, publicly listed pharmaceutical company founded in 1943. Its R&D is focused on dermatology and immunology-mediated skin diseases, with marketed products including treatments for psoriasis, atopic dermatitis, and actinic keratosis. The company has a strong tradition of academic collaboration in Catalonia and internationally.
B2SLab / IRIS-UPC contributes computational biology, machine learning, and bioinformatics expertise. The collaboration exemplifies the Catalan Industrial Doctorate programme’s goal of linking university research directly to industrial R&D needs.
Industrial Doctorate programme funded by the Generalitat de Catalunya (Secretaria d’Universitats i Recerca).