
Multilayer Perceptron and Learning
Advanced Topics in Machine Learning for Bioinformatics and Biomedical Engineering
March 9, 2026
MLPs and Backprop
Why backprop matters
- Modern bioinformatics models (CNNs for sequence motifs, transformers for proteins, autoencoders for single-cell) are trained by gradient-based optimization.
- Backpropagation is the efficient way to compute gradients of a loss with respect to millions of parameters.
Learning goals
- Understand backprop as the chain rule on a computational graph.
- Derive gradients for common layers and losses.
- Implement (and debug) backprop with sanity checks.
- Understand the why of activation functions.
Notation (we will reuse everywhere)
What a neuron/layer computes
A single unit (neuron) takes an input vector \(x \in \mathbb{R}^d\), computes:
- Linear step (logit / pre-activation)
\[ z = w^\top x + b \] - Output (activation)
\[ a = \phi(z) \]
For a typical layer
\[ z = Wx + b,\qquad h = \phi(z) \] where
Input vector: \(x \in \mathbb{R}^{d}\)
Parameters:
- weights \(W \in \mathbb{R}^{m \times d}\), maps \((d \to m)\)
- biases \(b \in \mathbb{R}^{m}\)
Hidden layer activation: \(h \in \mathbb{R}^{m}\)
Nonlinearity: \(\sigma(\cdot)\), or \(\phi(\cdot): \mathbb{R}^{m} \to \mathbb{R}^{m}\) (applied element-wise)
Output:
- logits \(z \in \mathbb{R}^{k}\)
- predictions \(\hat{y} \in \mathbb{R}^{k}\) (e.g., probabilities after softmax; for binary, often \(\hat{y}\in(0,1))\)
Loss: \(L \in \mathbb{R}\) (a scalar)
Notation (we will reuse everywhere)
For a typical layer
\[ z = Wx + b,\qquad h = \phi(z) \] where
d: input dimensionality = number of input features. Examples: gene-expression features, k-mer counts, embedding length.
m: hidden-layer width = number of neurons (units) in the hidden layer. This is a tunable architecture choice (a hyperparameter).
k: output dimensionality = number of outputs. Examples:
- multiclass classification: \(k\) = number of classes
- regression with \(k\) targets: \(k\) = number of predicted values
- binary classification: often \(k=1\) (single logit/probability), or sometimes (k=2) if using a 2-class softmax.
The big picture
A classical neural network
The big picture
Training loop:
Forward Pass: compute predictions \(\hat{y}\) from inputs \(x\) and parameters \(\theta\).
Loss Computation: measure mismatch \(L(\hat{y}, y)\). \(L\) measures how wrong a model’s prediction is on a given example.
Backward pass (backprop): compute \(\nabla_\theta L\) efficiently.
Update: \(\theta \leftarrow \theta - \eta \nabla_\theta L\) (or Adam, etc.).
About Loss functions
What is a loss function?
A loss function measures how wrong a model’s prediction is on a given example. Given: ¡
- input \(x\)
- target (ground truth) \(y\)
- model prediction \(\hat{y} = f_\theta(x)\)
the loss is a scalar: \[ L(\hat{y}, y) \in \mathbb{R} \]
We train the model by choosing parameters \(\theta\) that minimize the average loss over a dataset of \(n\) samples: \[ \min_{\theta}\; \frac{1}{n}\sum_{i=1}^{n} L\!\left(\hat{y}^{(i)}, y^{(i)}\right) \]
LSMS
The loss defines what good predictions mean.
Different tasks (regression, binary classification, multiclass) use different losses.
Overall loss functions
Loss functions are a big family. But in neural nets you can think of them in a few categories:
- Classification losses:
- Mainly
- Binary: sigmoid + BCE (log loss)
- Multiclass (single label): softmax + cross-entropy
- Multi-label: independent sigmoid + summed BCE
- Regression losses (continuous targets):
- e.g., MSE, MAE, Huber, log-cosh.
- Margin / ranking losses (separation or ordering):
- e.g., hinge loss, triplet loss, contrastive loss.
- Probabilistic / likelihood-based losses (negative log-likelihoods):
- e.g., Gaussian NLL, Poisson NLL (common for count data), categorical NLL.
Overall loss functions
- Distance / similarity losses (embeddings, metric learning):
- e.g., cosine loss, Euclidean distance objectives.
- Structured losses (sequence/structure outputs):
- e.g., CTC (speech-like sequence alignment), structured SVM losses (less common in basic NN courses).
- Regularization terms (often added to the loss):
- e.g., ( \(\lambda|W|_2^2\) ) (weight decay), ( \(\lambda|W|_1\) ). These aren’t “prediction losses” by themselves, but they modify the objective.
In practice, use just a few core ones: MSE (regression), BCE (binary/multi-label), and softmax cross-entropy (multiclass).
Regression loss functions
Mean squared error (MSE)
Used when targets are real-valued (e.g., expression level prediction, continuous phenotype).
For a single scalar prediction \(\hat{y}\in\mathbb{R}\): \[ L_{\text{MSE}}(\hat{y},y)=\frac{1}{2}(\hat{y}-y)^2 \] (The factor \(1/2\) is convenient: it cancels in derivatives.)
Gradient w.r.t. prediction: \[ \frac{\partial L_{\text{MSE}}}{\partial \hat{y}} = \hat{y}-y \]
When it works well: Gaussian noise / squared-error is acceptable, smooth optimization.
Sensitivity: strongly penalizes outliers.
Mean absolute error (MAE)
\[ L_{\text{MAE}}(\hat{y},y)=|\hat{y}-y| \] Subgradient: \[ \frac{\partial L_{\text{MAE}}}{\partial \hat{y}} \in \mathrm{sign}(\hat{y}-y) \] (Not differentiable at \(0\); optimizers use subgradients.)
When it works well: heavier-tailed noise, robust to outliers.
Tradeoff: gradient is constant (can be slower to converge near optimum).
Huber loss (smooth robust regression)
Interpolates between MSE (near 0) and MAE (far away). For residual \(r=\hat{y}-y\) and threshold \(\delta>0\): \[ L_{\delta}(r)= \begin{cases} \frac{1}{2}r^2 & |r|\le \delta \\ \delta(|r|-\frac{1}{2}\delta) & |r|>\delta \end{cases} \] Derivative: \[ \frac{\partial L_\delta}{\partial \hat{y}}= \begin{cases} r & |r|\le \delta \\ \delta\,\mathrm{sign}(r) & |r|>\delta \end{cases} \]
The Huber loss is the convolution of the absolute value function with the rectangular function, scaled and translated. Thus it “smoothens out” the former’s corner at the origin.
Practical: often a good default when outliers exist.
MSE vs MAE vs Huber

Binary classification losses
Sigmoid + binary cross-entropy (BCE)
Logit \(z\in\mathbb{R}\), probability \(\hat{y}=\sigma(z)\in(0,1)\), label \(y\in\{0,1\}\): \[ \hat{y}=\sigma(z)=\frac{1}{1+e^{-z}} \] \[ L_{\text{BCE}}(\hat{y},y)= -\Big(y\log\hat{y} + (1-y)\log(1-\hat{y})\Big) \]
A key simplification for backprop is the derivative w.r.t. the logit1: \[ \frac{\partial L_{\text{BCE}}}{\partial z}=\hat{y}-y \]
BCE with logits (numerically stable)
In implementations you typically avoid computing \(\log(1-\sigma(z))\) directly. A stable form is: \[ L_{\text{BCE-logits}}(z,y)=\log\big(1+e^{z}\big) - yz \] (which is equivalent to BCE after algebra).
Why it matters: prevents overflow/underflow for large \(|z|\).
BCE as a function of logit

Multiclass classification losses
Softmax + categorical cross-entropy
Logits \(z\in\mathbb{R}^{k}\), probabilities \(\hat{y}\in\Delta^{k-1}\) 1: \[ \hat{y}_i = \mathrm{softmax}(z)_i=\frac{e^{z_i}}{\sum_{j=1}^k e^{z_j}} \]
For one-hot \(y\in\{0,1\}^k\): \[ L_{\text{CE}}(\hat{y},y)= -\sum_{i=1}^k y_i \log(\hat{y}_i) \]
Softmax + categorical cross-entropy
Backprop simplification: \[ \frac{\partial L_{\text{CE}}}{\partial z}=\hat{y}-y \]
It is the multiclass analogue of BCE+sigmoid.
Stable computation (log-sum-exp trick)
Use: \[ \log\sum_j e^{z_j} = \alpha + \log\sum_j e^{z_j-\alpha},\quad \alpha=\max_j z_j \] to avoid overflow when some logits are large.
Cross-entropy vs margin for the correct class

Multi-label classification (independent labels)
e.g. predicting multiple functional GO annotations per protein (several labels can be 1 simultaneously).
Use \(k\) independent logits \(z\in\mathbb{R}^k\) and apply sigmoid per label: \[ \hat{y}_j = \sigma(z_j) \] Loss is a sum (or mean) of BCE across labels: \[ L=\sum_{j=1}^k \mathrm{BCE}(\hat{y}_j, y_j) \]
Gradient still has a simple per-label form: \[ \frac{\partial L}{\partial z_j} = \hat{y}_j - y_j \]
Handling class imbalance
Bioinformatics datasets often have strong imbalance (rare motifs, rare cell types, rare variants).
Weighted BCE
Let positive weight \(w_+>0\) and negative weight \(w_->0\): \[ L = -\left(w_+\,y\log\hat{y} + w_-\,(1-y)\log(1-\hat{y})\right) \] This changes the gradient scale so minority classes influence training more.
Focal loss (down-weights easy examples)
Often used for extreme imbalance. Define \(p_t=\hat{y}\) if \(y=1\), else \(p_t=1-\hat{y}\): \[ L_{\text{focal}} = -(1-p_t)^\gamma \log(p_t) \] - \(\gamma>0\) focuses learning on hard cases (where \(p_t\) is small). - Often combined with class weights.
Intuition: if an example is already confidently correct, it contributes less to the loss.
BCE vs focal loss (binary)

Ranking / margin-based losses (brief)
Some tasks are naturally ranking problems (e.g., scoring candidate interactions).
Hinge loss (binary, margin)
With label \(y\in\{-1,+1\}\) and score \(s\in\mathbb{R}\): \[ L_{\text{hinge}}(s,y)=\max(0, 1 - y s) \] Encourages \(ys\ge 1\) (a margin).
Often used in SVMs; less common than cross-entropy in modern deep nets, but useful conceptually.
Choosing a loss: practical guidance (bioinformatics)
- Regression of continuous targets: start with MSE; switch to Huber if outliers.
- Binary classification: BCE with logits (stable), consider class weights for imbalance.
- Multiclass (one label per sample): softmax + cross-entropy.
- Multi-label: sigmoid per label + summed BCE.
- Extreme imbalance / detection-like tasks: consider focal loss (plus weights).
- Uncertainty/probabilistic modeling: cross-entropy is a negative log-likelihood under a categorical model; MSE corresponds to a Gaussian likelihood (fixed variance).
Backpropagation
The big picture
If \(\theta\) is the collection of all parameters, say \(\theta = (\theta_1,\theta_2,\dots,\theta_n)\) then the gradient is the vector of partial derivatives: \(\nabla_\theta L = \left( \frac{\partial L}{\partial \theta_1}, \frac{\partial L}{\partial \theta_2}, \dots, \frac{\partial L}{\partial \theta_n} \right)\)
To train a network we will do:
- Forward Pass: compute predictions \(\hat{y}\) from inputs \(x\) and parameters \(\theta\).
- Loss Computation: measure mismatch \(L(\hat{y}, y)\).
- Backward pass (backprop): compute \(\nabla_\theta L\) efficiently.
- Update: \(\theta \leftarrow \theta - \eta \nabla_\theta L\) (or Adam, etc.). So \(\theta_j \leftarrow \theta_j - \eta \frac{\partial L}{\partial \theta_j} \quad \text{for each } j\)
Example single neuron with sigmoid
\[ a = w^\top x + b,\qquad \hat{y}=\sigma(a)=\frac{1}{1+e^{-a}} \]
Binary cross-entropy (BCE) for label \(y\in\{0,1\}\): \[ L(\hat{y},y) = -\left(y\log \hat{y} + (1-y)\log(1-\hat{y})\right) \]
We want gradients: \(\frac{\partial L}{\partial w}\) and \(\frac{\partial L}{\partial b}\).
Computational graph viewpoint
flowchart LR x((x)) --> a[$$a = w \cdot x + b$$] w((w)) --> a b((b)) --> a a --> yhat["$$\hat y = \sigma (a)$$"] yhat --> L["$$L(\hat y, y)$$"] y((y)) --> L
Chain rule in one line
Backprop = applying the chain rule along the edges of a graph. \[ a = w^\top x + b,\qquad \hat{y}=\sigma(a)=\frac{1}{1+e^{-a}} \]
\[ L(\hat{y},y) = -\left(y\log \hat{y} + (1-y)\log(1-\hat{y})\right) \]
If \(L\) depends on \(\hat{y}\), which depends on \(a\), which depends on \(w\):
The problem
\[ \frac{\partial L}{\partial w} = \frac{\partial L}{\partial \hat{y}} \cdot \frac{\partial \hat{y}}{\partial a} \cdot \frac{\partial a}{\partial w} \]
Backprop organizes this so we reuse intermediate derivatives.

Backpropagation
The problem
\[ \frac{\partial L}{\partial w} = \frac{\partial L}{\partial \hat{y}} \cdot \frac{\partial \hat{y}}{\partial a} \cdot \frac{\partial a}{\partial w} \]
and then
\[ w \leftarrow w - \eta \frac{\partial L}{\partial w} \]
For our example
Sigmoid: \[ \sigma'(a)=\sigma(a)\left(1-\sigma(a)\right) \]
Affine: \[ a = w^\top x + b \quad\Rightarrow\quad \frac{\partial a}{\partial w}=x,\; \frac{\partial a}{\partial b}=1 \]
Let’s derive \(\partial L/\partial a\)
Let’s derive \(\frac{\partial L}{\partial a}\)
Binary classifier (single neuron):
\[ a = w^\top x + b,\qquad \hat{y}=\sigma(a)=\frac{1}{1+e^{-a}} \]
Binary cross-entropy (BCE):
\[ L(\hat{y},y)= -\Big(y\log \hat{y} + (1-y)\log(1-\hat{y})\Big), \quad y\in\{0,1\} \]
We want the gradient w.r.t. the logit \(a\): \(\frac{\partial L}{\partial a}\).
Step 1: compute \(\frac{\partial L}{\partial \hat{y}}\)
Differentiate BCE w.r.t. \(\hat{y}\):
\[ \frac{\partial L}{\partial \hat{y}} = -\left( y\cdot \frac{1}{\hat{y}} + (1-y)\cdot \frac{-1}{1-\hat{y}} \right) = -\frac{y}{\hat{y}}+\frac{1-y}{1-\hat{y}} \]
Put over a common denominator:
\[ \frac{\partial L}{\partial \hat{y}} = \frac{-y(1-\hat{y})+(1-y)\hat{y}}{\hat{y}(1-\hat{y})} = \frac{\hat{y}-y}{\hat{y}(1-\hat{y})} \]
Step 2: compute \(\displaystyle \frac{\partial \hat{y}}{\partial a}\)
Sigmoid derivative:
\[ \hat{y}=\sigma(a)=\frac{1}{1+e^{-a}} \quad\Rightarrow\quad \frac{\partial \hat{y}}{\partial a} = \sigma(a)(1-\sigma(a)) = \hat{y}(1-\hat{y}) \]
Step 3: chain rule and the simplification
By the chain rule:
\[ \frac{\partial L}{\partial a} = \frac{\partial L}{\partial \hat{y}} \cdot \frac{\partial \hat{y}}{\partial a} \]
Substitute the two results:
\[ \frac{\partial L}{\partial a} = \left(\frac{\hat{y}-y}{\hat{y}(1-\hat{y})}\right) \cdot \left(\hat{y}(1-\hat{y})\right) = \hat{y}-y \]
The simplification !
This cancellation is why sigmoid + BCE is so clean in backprop:
the gradient at the logit is just prediction minus label!!.
For the parameters \(w,b\)
Since \(a=w^\top x + b\):
\[ \frac{\partial a}{\partial w}=x,\qquad \frac{\partial a}{\partial b}=1 \]
Therefore:
\[ \frac{\partial L}{\partial w} = \frac{\partial L}{\partial a}\frac{\partial a}{\partial w} = (\hat{y}-y)x, \qquad \frac{\partial L}{\partial b} = \hat{y}-y \]
Same pattern for softmax + cross-entropy
For multiclass logits \(z\in\mathbb{R}^k\), \(\hat{y}=\mathrm{softmax}(z)\), and one-hot \(y\):
\[ L = -\sum_{i=1}^k y_i\log(\hat{y}_i) \]
A key result (analogous to BCE+sigmoid) is:
\[ \frac{\partial L}{\partial z}=\hat{y}-y \]
So in both cases, the error signal at the final logits is \(\hat{y}-y\).
(extra) For a Vectorized setup (batch + layer view)
Batch of size \(n\):
- Inputs \(X \in \mathbb{R}^{n\times d}\)
- Weights \(W \in \mathbb{R}^{1\times d}\), bias \(b\in\mathbb{R}\) (binary case, one logit)
- Logits \(a \in \mathbb{R}^{n\times 1}\)
- Probabilities \(\hat{Y}\in(0,1)^{n\times 1}\)
- Labels \(Y\in\{0,1\}^{n\times 1}\)
Layer view: \[ a = XW^\top + b \qquad \hat{Y} = \sigma(a) \]
We use the mean BCE over the batch: \[ L = \frac{1}{n}\sum_{i=1}^n \ell(\hat{y}^{(i)},y^{(i)}) \]
Binary cross-entropy per sample
For sample \(i\): \[ \ell(\hat{y}^{(i)},y^{(i)})= -\Big(y^{(i)}\log \hat{y}^{(i)} + (1-y^{(i)})\log(1-\hat{y}^{(i)})\Big) \]
Derivative wrt \(\hat{y}^{(i)}\): \[ \frac{\partial \ell}{\partial \hat{y}^{(i)}} = \frac{\hat{y}^{(i)}-y^{(i)}}{\hat{y}^{(i)}(1-\hat{y}^{(i)})} \]
Sigmoid derivative: \[ \frac{\partial \hat{y}^{(i)}}{\partial a^{(i)}}=\hat{y}^{(i)}(1-\hat{y}^{(i)}) \]
Error signal at the logits (vector form)
Chain rule per sample: \[ \frac{\partial \ell}{\partial a^{(i)}} = \frac{\partial \ell}{\partial \hat{y}^{(i)}} \cdot \frac{\partial \hat{y}^{(i)}}{\partial a^{(i)}} = \hat{y}^{(i)}-y^{(i)} \]
Stacking all samples: \[ \frac{\partial \ell}{\partial a} = \hat{Y}-Y \quad \in \mathbb{R}^{n\times 1} \]
For the mean loss: \[ \boxed{ \frac{\partial L}{\partial a} = \frac{1}{n}(\hat{Y}-Y) } \]
Note
You will often see \(\delta := \frac{\partial L}{\partial a}\) called the error signal. With mean reduction, it carries the \(\tfrac{1}{n}\) factor.
Gradients for the layer parameters \(W,b\)
Recall: \[ a = XW^\top + b \]
Using matrix calculus:
- Since \(a\) depends linearly on \(W\): \[ \boxed{ \frac{\partial L}{\partial W} = \left(\frac{\partial L}{\partial a}\right)^\top X } \quad\Rightarrow\quad \frac{\partial L}{\partial W}\in\mathbb{R}^{1\times d} \]
Gradients for the layer parameters \(W,b\)
- Bias gradient (sum over batch): \[ \boxed{ \frac{\partial L}{\partial b} = \sum_{i=1}^n \frac{\partial L}{\partial a^{(i)}} } \quad\Rightarrow\quad \frac{\partial L}{\partial b}\in\mathbb{R} \]
With mean loss: \[ \frac{\partial L}{\partial W}=\frac{1}{n}(\hat{Y}-Y)^\top X, \qquad \frac{\partial L}{\partial b}=\frac{1}{n}\sum_{i=1}^n(\hat{y}^{(i)}-y^{(i)}) \]
Extension: \(k\)-class softmax + cross-entropy (batch)
Now logits \(Z\in\mathbb{R}^{n\times k}\), probabilities \(\hat{Y}\in\mathbb{R}^{n\times k}\), one-hot targets \(Y\in\{0,1\}^{n\times k}\).
\[ \hat{Y}=\mathrm{softmax}(Z) \quad (\text{row-wise}) \] \[ L = -\frac{1}{n}\sum_{i=1}^n \sum_{c=1}^k Y_{ic}\log(\hat{Y}_{ic}) \]
Key result (same pattern): \[ \boxed{ \frac{\partial L}{\partial Z}=\frac{1}{n}(\hat{Y}-Y) } \]
So the last-layer error signal is still prediction minus target.
Where this plugs into an MLP
For an MLP ending in logits \(Z\):
- Compute \(\hat{Y}\) (sigmoid for binary, softmax for multiclass)
- Compute \(\delta^{(\text{out})} = \frac{\partial L}{\partial Z}\)
Then backprop into the previous layer: \[ \delta^{(\text{prev})} = \left(\delta^{(\text{out})}W^{(\text{out})}\right)\odot \phi'(A^{(\text{prev})}) \]
This is the standard backprop recursion.
BCE + sigmoid and the magic simplification
For BCE with sigmoid output, the gradient w.r.t. the pre-activation \(a\) is:
\[ \frac{\partial L}{\partial a} = \hat{y} - y \]
Then \[ \frac{\partial L}{\partial w} = (\hat{y}-y)\,x,\qquad \frac{\partial L}{\partial b} = (\hat{y}-y) \]
Tip
This is why logistic regression and binary classifiers are so clean to implement.
Visual intuition: derivatives of common activations


Why backprop is efficient
Naively computing each partial derivative separately is expensive.
Backprop computes all parameter gradients in time proportional to: - one forward pass + one backward pass
Roughly:
- \(O(\#\text{edges in graph})\),
- not \(O(\#\text{parameters} \times \#\text{operations})\).
Backprop as “messages” (reverse-mode autodiff)
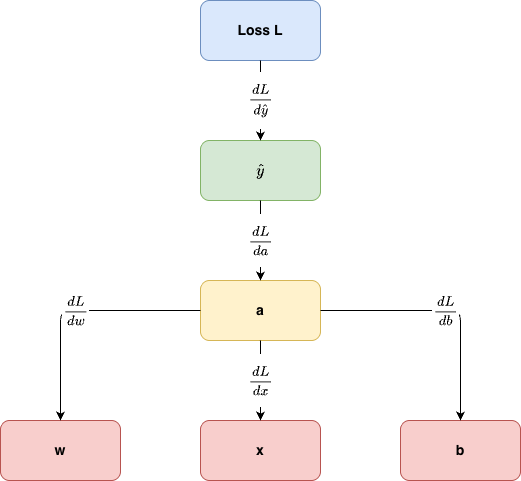
During the backward pass each node receives an upstream gradient \[ \bar{v} := \frac{\partial L}{\partial v} \] and sends downstream gradients to its parents.
What a neuron/layer computes
A single unit (neuron) takes an input vector \(x \in \mathbb{R}^d\), computes:
- Linear step (logit / pre-activation)
\[ z = w^\top x + b \] - Output (activation)
\[ a = \phi(z) \]
Layer view (many neurons at once):
\[ z = Wx + b,\qquad a = \phi(z) \] where \(\phi\) is applied element-wise for most common activations.
Role of Activation Functions
- Introduce non-linearity into perceptron output.
- Model the behavior of biological neurons.
- Examples: Step function, Sigmoid, ReLU.
Why activation functions?
A neural network layer typically computes:
\[ \mathbf{z} = \mathbf{W}\mathbf{x} + \mathbf{b}, \qquad \mathbf{a} = \phi(\mathbf{z}) \]
Without a nonlinear activation \(\phi(\cdot)\), stacking layers collapses into a single linear transformation, so the network cannot represent nonlinear relationships (crucial in bioinformatics: regulatory interactions, epistasis, nonlinear signal-response curves, etc.).
Activation functions mainly affect:
- Expressivity (nonlinearity)
- Optimization (gradient flow, saturation, dead units)
- Output interpretation (probabilities vs real values)
Summary table
| Activation | Typical use | Range | Key risk |
|---|---|---|---|
| Identity (linear) | Regression outputs | \((-\infty,\infty)\) | No nonlinearity |
| Sigmoid | Binary probability outputs | \((0,1)\) | Saturation, vanishing gradients |
| Tanh | Hidden layers (older), RNNs | \((-1,1)\) | Saturation, vanishing gradients |
| ReLU | Default hidden layers | \([0,\infty)\) | “Dying ReLU” (zero gradient) |
| Leaky ReLU | Hidden layers | \((-\infty,\infty)\) | Slope choice |
| ELU | Hidden layers | \((-\alpha,\infty)\) | Slightly costlier than ReLU |
| Softplus | Smooth ReLU alternative | \((0,\infty)\) | Still saturates for very negative inputs |
| GELU | Transformers / modern deep nets | \((-\infty,\infty)\) | More compute than ReLU |
| Softmax | Multi-class probability outputs | \((0,1)\) with sum 1 | Numerical stability if naive |
Identity (Linear)
\[ \phi(x) = x \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \(\mathbb{R}\)
- Differentiable everywhere; \(\phi'(x)=1\)
- Monotone increasing, no saturation
- Used mainly as the final layer for regression (e.g., predicting expression levels as real values)

Sigmoid (Logistic)
\[ \sigma(x) = \frac{1}{1 + e^{-x}} \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \((0,1)\) (interpretable as a probability)
- Smooth and differentiable everywhere
- Derivative: \[ \sigma'(x) = \sigma(x)\,[1-\sigma(x)] \]
- Saturates for large \(|x|\) \(\Rightarrow\) vanishing gradients
- Commonly used for binary classification outputs (e.g., disease vs control)

Hyperbolic Tangent (tanh)
\[ \tanh(x) = \frac{e^x - e^{-x}}{e^x + e^{-x}} \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \((-1,1)\) (zero-centered outputs often help optimization vs sigmoid)
- Smooth and differentiable everywhere
- Derivative: \[ \frac{d}{dx}\tanh(x) = 1 - \tanh^2(x) \]
- Saturates for large \(|x|\) \(\Rightarrow\) vanishing gradients
- Historically common in hidden layers and RNNs; now often replaced by ReLU-family activations

ReLU (Rectified Linear Unit)
\[ \mathrm{ReLU}(x) = \max(0, x) \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \([0,\infty)\)
- Piecewise linear; derivative: \[ \mathrm{ReLU}'(x)= \begin{cases} 0 & x < 0 \\ 1 & x > 0 \end{cases} \] (At \(x=0\) it is not differentiable; in practice, a subgradient is used.)
- fast, encourages sparse activations, mitigates vanishing gradients on \(x>0\)
- Dying ReLU — if a unit stays at \(x<0\), its gradient is 0 and it may stop learning

Leaky ReLU
\[ \phi(x)= \begin{cases} x & x\ge 0 \\ \alpha x & x<0 \end{cases} \quad \text{with } \alpha \in (0,1) \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \(\mathbb{R}\)
- Piecewise linear, not differentiable at 0 (handled with subgradients)
- Derivative: \[ \phi'(x)= \begin{cases} 1 & x>0 \\ \alpha & x<0 \end{cases} \]
- Reduces dying ReLU by allowing a small gradient when \(x<0\)

ELU (Exponential Linear Unit)
\[ \mathrm{ELU}(x)= \begin{cases} x & x\ge 0 \\ \alpha\left(e^x - 1\right) & x<0 \end{cases} \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \((-\alpha,\infty)\)
- Smooth on \(x<0\), continuous everywhere; often chosen \(\alpha=1\)
- Derivative: \[ \mathrm{ELU}'(x)= \begin{cases} 1 & x>0 \\ \alpha e^x & x<0 \end{cases} \]
- Negative outputs can help keep activations closer to zero mean (sometimes improving optimization)

Softplus (Smooth ReLU)
\[ \mathrm{Softplus}(x)=\ln\left(1+e^x\right) \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \((0,\infty)\)
- Smooth approximation to ReLU
- Derivative: \[ \frac{d}{dx}\mathrm{Softplus}(x)=\sigma(x) \]
- For very negative \(x\), gradients become small (still some saturation), but it avoids the hard link at 0

GELU (Gaussian Error Linear Unit)
A common approximation used in practice is:
\[ \mathrm{GELU}(x) \approx \frac{1}{2}x\left[1+\tanh\left(\sqrt{\frac{2}{\pi}}(x+0.044715x^3)\right)\right] \]
Properties
- Domain: \(\mathbb{R}\)
- Range: \(\mathbb{R}\)
- Smooth, non-monotonic near the origin (it can slightly down-weight small negative inputs)
- Common in Transformers and other modern architectures
- More computationally expensive than ReLU/Leaky ReLU

Softmax (Multi-class output)
For logits \(\mathbf{z} \in \mathbb{R}^K\),
\[ \mathrm{Softmax}(z_i)=\frac{e^{z_i}}{\sum_{j=1}^{K} e^{z_j}} \]
Properties
- Domain: \(\mathbb{R}^K\)
- Range: \((0,1)^K\) with \(\sum_i p_i = 1\)
- Outputs are probabilities: each in \((0,1)\) and \(\sum_i p_i = 1\)
- Invariant to adding a constant: \[ \mathrm{Softmax}(\mathbf{z})=\mathrm{Softmax}(\mathbf{z}+c) \]
- Needs numerical stability: compute with \(\mathbf{z}-\max(\mathbf{z})\)
- Typical use: multi-class classification output layer

All of them


Learning Curves
What is a Learning Curve?
Definition
A learning curve shows a score of an estimator for a varying number of training samples (or epochs).
Intuitively, a model should increase its learning (score) with more experience (samples or epochs).
It is common practice to plot dual curves simultaneously:
- the training dataset curve, and
- the validation dataset curve.
What does a Learning Curve answer?
- How much does the model benefit from adding more training data?
- Does the estimator suffer more from variance error or bias error?
Learning curves are heavily used when training deep neural networks, where each epoch provides a natural evaluation point.
Setup: what to track
You can configure your learning curve to maximise or minimise a score:
- Maximise (higher is better)
- e.g., classification accuracy, F1-score, AUROC
- Minimise (lower is better, target \(\approx\) 0)
- e.g., loss (MSE, cross-entropy, RMSE) — the most common choice in deep learning
During training, at each step you evaluate two quantities:
- Train LC
- Score on the training set — measures how well the model is learning.
- Val LC
- Score on a held-out validation set — measures how well the model is generalising.
Diagnosing Model Behaviour
What can we learn?
From the dynamics and shape of the learning curves we can diagnose:
About our model:
- Underfitting
- Good fitting
- Overfitting
About our data:
- Unrepresentative training dataset
- Unrepresentative validation dataset (handle with care)
These patterns hint at issues with model capacity, dataset size, and generalisation.
Underfit — model too simple
Underfit condition
The model cannot obtain a sufficiently low error value on the training set.

Flat curves at high loss
The model does not have sufficient capacity for the complexity of the dataset.
Causes:
- Model too simple (too few parameters / layers)
- Features not expressive enough
- Learning rate too high
Remedy: increase model capacity, add layers, or improve features.
Underfit — training stopped too early
Underfit condition
The model cannot obtain a sufficiently low error value on the training set.

Dec. Train loss · Dec. Val loss
Both curves are still falling — the model has capacity to keep improving but training was stopped prematurely.
Causes:
- Too few epochs
- Very aggressive early stopping criterion
Remedy: train for more epochs; revisit early-stopping patience.
Overfit
Overfit condition
The model learns the training data too well, at the cost of increased generalisation error.

Dec. Train loss · Inc. Val loss
Training loss keeps decreasing while validation loss starts to increase — a clear sign the model is memorising rather than learning.
Remedies:
- Regularisation (L2, dropout)
- Data augmentation
- Reduce model capacity
- Early stopping at the divergence point
Good Fit
Good fit condition
The model learns the training data correctly and generalises well.
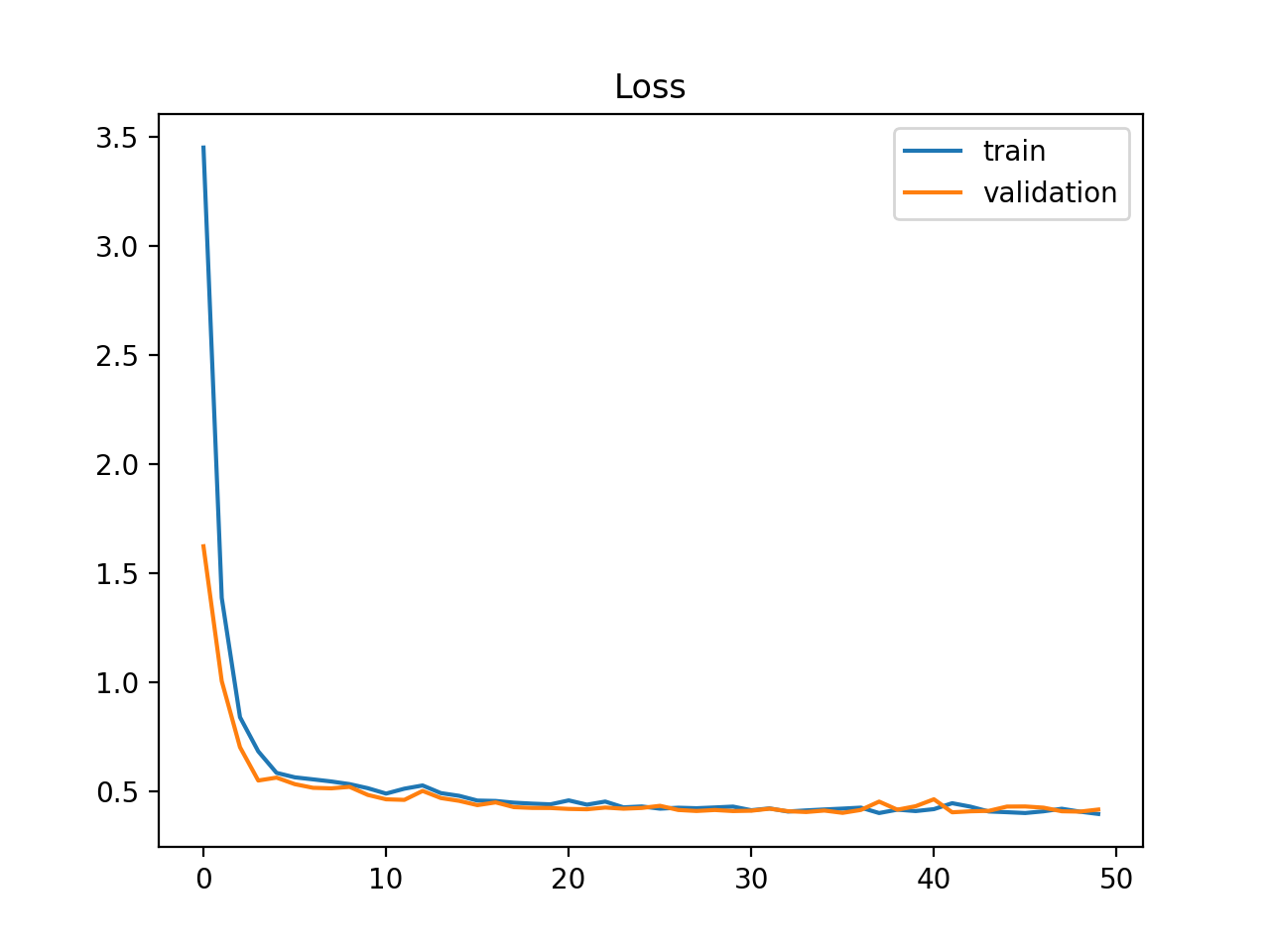
Stable Train loss · Stable Val loss
- Training loss decreases and stabilises \(\checkmark\)
- Validation loss follows a similar path \(\checkmark\)
- Both reach a similar final value \(\checkmark\)
A small, stable gap between train and val loss is acceptable — a perfect gap of zero is unlikely in practice.
Unrepresentative Training Set
Unrepresentative training condition
The training data does not provide sufficient information to explain the validation distribution. (Sample size? Distribution shift?)

Dec. Train loss · Stable Val loss · Growing gap
- Training loss decreases steadily
- Validation loss stabilises at a much higher value
- The gap between the two curves widens over time
Remedies:
- Collect more training data
- Apply data augmentation
- Check for train/val distribution mismatch
Unrepresentative Validation Set — noisy
Unrepresentative validation condition
The validation data may not provide sufficient information to evaluate generalisation reliably.

Stable Train loss · Noisy Val loss
- Training loss looks like a good or acceptable fit
- Validation loss shows high variance — erratic behaviour with no clear trend
Causes:
- Validation set too small
- Validation set not stratified / shuffled
Remedy: increase validation set size; use k-fold cross-validation.
Unrepresentative Validation Set — val below train
Unrepresentative validation condition
Validation loss systematically lower than training loss — often an artefact of the training procedure.

Stable Train loss · Lower Val loss · Wide gap
- Training loss looks like a good fit
- Validation loss converges to a value lower than training loss
- The gap is persistent and wide
Common cause: dropout or other stochastic regularisation active during training but disabled during evaluation, making val loss artificially lower.
Not always a problem — understand the cause first.
Practical Notes
Why overfitting matters in bioinformatics (and everywhere)
A deep model can look brilliant on the training set and still be useless on new samples.
Think about a classifier trained to distinguish tumour subtypes from gene-expression profiles. If the network learns quirks of the training cohort rather than stable biological patterns, it will not generalise to patients from another lab, hospital, or sequencing batch.
\[ \mathcal{L}_{\text{train}} \downarrow \quad \text{while} \quad \mathcal{L}_{\text{val}} \uparrow \]
\[ \text{generalisation gap} = \mathcal{L}_{\text{val}} - \mathcal{L}_{\text{train}} \]
Note
A widening gap between training and validation loss is the classic warning sign: the model is remembering the data, not learning the signal.
Tip
In biology this is especially dangerous — datasets are often small, noisy, high-dimensional, and heterogeneous.

A healthy model improves on both sets; an overfit model keeps improving only on training data.
Recap
NN
Dropout
About dropout
Dropout
Dropout randomly switches off neurons during training. The network cannot rely on one convenient pathway, so it learns more robust, distributed representations.
\[ \tilde{h}_i = h_i \cdot m_i, \qquad m_i \sim \mathrm{Bernoulli}(1-p) \]
At test time, dropout is disabled and activations are rescaled by \((1-p)\) to preserve expected magnitude.
- Too little \((p \approx 0.1)\): barely any effect
- Too much \((p > 0.7)\): optimisation becomes difficult
- Typical range: 0.2–0.5 on dense layers
Tip
For omics tabular data, dropout is most effective in the fully connected layers — not at the raw input, where every feature may already be scarce.
The pattern changes every batch, forcing the network to learn redundant representations.
Weight penalties: L1 and L2 regularisation
L2 (weight decay)
\[ \mathcal{L} = \mathcal{L}_{\text{data}} + \lambda \sum_i w_i^2 \]
Large weights are penalised quadratically. The model keeps all features but with modest coefficients.
L1 (Lasso)
\[ \mathcal{L} = \mathcal{L}_{\text{data}} + \lambda \sum_i |w_i| \]
Promotes sparsity — some weights shrink to exactly zero, acting like automatic feature selection.
Warning
\(\lambda\) controls regularisation strength. Too small → no effect. Too large → underfitting. Tune it on validation data.
Tip
In transcriptomics or proteomics, L1 is attractive because sparse solutions are easier to interpret biologically.
L2 discourages extreme weights; L1 pushes many weights to exactly zero — automatic feature selection.
Batch normalisation and label smoothing
Batch normalisation
Batch norm standardises each layer’s output during training:
\[ \hat{x}_i = \frac{x_i - \mu_{\mathcal{B}}}{\sqrt{\sigma^2_{\mathcal{B}} + \varepsilon}}, \qquad y_i = \gamma \hat{x}_i + \beta \]
where \(\mu_{\mathcal{B}}, \sigma^2_{\mathcal{B}}\) are computed over the current mini-batch and \(\gamma, \beta\) are learnable. The mini-batch dependence also injects a small, healthy amount of noise.
Label smoothing
Instead of hard targets \((0, 1, 0)\), soften them slightly:
\[ y^{\text{smooth}} = (1-\alpha)\,y + \frac{\alpha}{K} \]
This reduces overconfidence and improves calibration.
Note
Label smoothing is particularly useful in biological classification tasks where labels are noisy or borderline cases exist.
Batch norm stabilises the learning signal; label smoothing prevents the model from becoming overconfident.
Early stopping and data augmentation
Early stopping
Monitor validation loss. Stop when it has not improved for \(k\) consecutive epochs (patience):
\[ \text{stop if } \mathcal{L}_{\text{val}}^{(t)} > \min_{t' < t} \mathcal{L}_{\text{val}}^{(t')} \;\text{for}\; k \text{ steps} \]
No extra parameters. Save the best checkpoint and restore it.
Data augmentation
Create extra training examples with label-preserving transforms:
| Data type | Example transforms |
|---|---|
| Microscopy / histology | flip, rotate, crop, colour jitter |
| DNA sequences | reverse complement, masking |
| Protein sequences | token masking, substitutions |
| Expression matrices | mild noise after normalisation |
Warning
Augmentation must preserve the label. A transform that changes the biology is not regularisation — it is corruption.
Early stopping controls training time; augmentation increases effective dataset diversity.
A sensible recipe in practice
Start simple
Use a model no larger than necessary. In small biological datasets, architecture size matters as much as clever regularisation.
Combine techniques
A common recipe is weight decay + early stopping, with dropout added if the dense layers are clearly overfitting.
Validate honestly
Split by patient, experiment, or batch — not by random rows. In bioinformatics, leakage hides in the structure of the data.
Practical checklist
- Watch both training and validation curves throughout training.
- Tune \(\lambda\), dropout rate, and patience on validation data — never on the test set.
- Prefer biologically plausible augmentation strategies.
- Be suspicious of near-perfect training accuracy on a small dataset.
- Interpret success in terms of generalisation, not memorisation.
Note
The most important idea is not a specific trick. It is the habit of asking: “Will this model still work on new biological samples?”

b2slab.upc.edu alexandre.perera@upc.edu