Does your mouse model actually look like a human disease? Now there’s a tool to check, cell by cell
The problem with animal models
Before a drug reaches clinical trials, it must work in an animal model. But here lies one of the most persistent frustrations in biomedical research: a treatment that cures mice often fails in humans. Part of the reason is that we rarely have a rigorous, quantitative answer to the question: how similar is this mouse model to the human disease, at the biological level?
Most comparisons are qualitative — researchers rely on shared symptoms or broad gene expression overlap. What has been missing is a method that works at the resolution of individual cell types, individual pathways, and individual genes, and that produces a number: a similarity score.
What singIST does
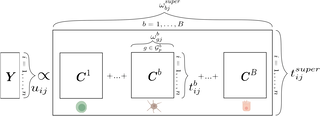
Our new method, singIST (single-cell Integrative Similarity Tool), addresses this directly. Published in PLOS Computational Biology, singIST takes single-cell RNA sequencing (scRNA-seq) data from both a disease model and human patients and computes how well the model captures the human disease at three nested levels:
- Pathway level — are the same biological processes disrupted?
- Cell type level — are the same cell populations involved?
- Gene level — are the same individual genes differentially expressed?
Critically, singIST accounts for two biological realities that simpler methods ignore: gene conservation between species (not every human gene has a functional mouse equivalent), and differences in cell type composition between species (a cell type present in human skin may be rare or absent in mouse skin).
The method produces interpretable similarity scores at each level, making it possible to say not just “this model is broadly similar” but “this model captures keratinocyte biology well but misses the dendritic cell response entirely”.
Testing on skin disease models
We validated singIST on three mouse models of atopic dermatitis (AD), a common inflammatory skin disease, and additionally applied it to human hidradenitis suppurativa (HS) data. These diseases share some features but differ mechanistically, making them a good test of the method’s discriminative power.
singIST reproduced established knowledge — for instance, correctly identifying which mouse model best recapitulates the Th2-skewed immune response characteristic of human AD — while also generating new hypotheses about specific pathways and cell types that each model captures or misses.
Why it matters for drug development
Choosing the wrong preclinical model is a major driver of clinical trial failure. singIST offers a systematic, reproducible way to audit candidate models before committing to expensive and time-consuming experiments. Rather than discovering post-hoc that a model missed a key pathway, researchers can now make that comparison up front.
The tool is freely available and designed to work with standard scRNA-seq pipelines, meaning it can be integrated into existing workflows without requiring additional data collection.
This work was led by Aitor Moruno-Cuenca in collaboration with Francesc Fernández-Albert and colleagues at B2SLab (IRIS-UPC) and the University of Michigan.
The paper is available at: Moruno-Cuenca A, Picart-Armada S, Bogle R, Fox J, Tsoi LC, Gudjonsson JE, Perera-Lluna A, Fernández-Albert F. singIST: An integrative method for comparative single-cell transcriptomics between disease models and humans. PLOS Computational Biology, 2026. https://doi.org/10.1371/journal.pcbi.1014002