Teaching a machine to smell: can an electronic nose tell good olive oil from bad?
The electronic nose: a sensor with an identity crisis
The idea is appealing: a device that sniffs a sample and tells you something useful about it — whether food is fresh, whether air is clean, whether a patient has a certain disease. This is the promise of the electronic nose (e-nose), an array of gas sensors whose combined response encodes chemical information about a vapour.
For decades, e-noses worked by analogy with the human nose: an untargeted, pattern-based approach where the overall sensor response was matched to known samples, without needing to identify individual molecules. More recently, researchers have started incorporating analytical instruments — such as ion mobility spectrometers (IMS) or gas chromatographs — to add chemical specificity. These instrument-based e-noses can distinguish individual compounds, not just broad odour profiles.
But this added specificity comes with a cost: more data, more complexity, and a greater risk of building models that memorise training examples rather than generalising to new samples.
Preprocessing before modelling: a neglected step
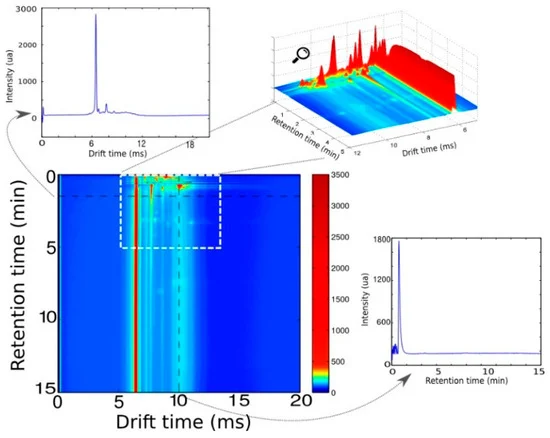
In a study published in Sensors, our group introduced a multivariate signal processing workflow specifically designed for datasets from instrument-based e-noses — validated on a multi-capillary column IMS (MCC-IMS) platform.
The key insight is that instrument-based e-noses, despite their greater specificity, still produce data that requires correction before reliable models can be built. Baseline drift, retention time variability, and instrument noise can all confound the signal. Traditional approaches handle this through peak integration — identifying and quantifying individual compounds — but this requires expert knowledge, is labour-intensive, and defeats the purpose of an untargeted screening device.
Our workflow instead applies preprocessing in an untargeted way: no compound identification required, no manual peak picking. The approach treats the full sensor signal as a multivariate time series and applies corrections that suppress systematic noise while preserving discriminative information.
The result: simpler models that generalise better
The core finding is that preprocessing reduces overfitting and produces more parsimonious models — models that achieve the same or better classification accuracy with fewer variables. This matters practically: a simpler model is more robust when applied to new samples, and easier to interpret and validate.
We demonstrated this on an olive oil quality control dataset, where the task was to classify olive oils by quality grade from IMS sensor data. After applying the preprocessing workflow, classification accuracy was maintained while model complexity dropped substantially. The models built on preprocessed data generalised better to held-out samples than those built on raw data.
Beyond olive oil
Olive oil quality control is a useful test case — economically important (adulteration is a significant problem in the olive oil market) and technically demanding (the chemical differences between quality grades are subtle). But the workflow is generic: it applies to any MCC-IMS dataset, and more broadly to any instrument-based e-nose platform producing multivariate spectral data.
Potential applications extend to food authenticity testing, environmental monitoring, and eventually clinical breathomics — the analysis of exhaled breath for disease markers.
The paper is available at: Fernandez L, Oller-Moreno S, Fonollosa J, Garrido-Delgado R, Arce L, Martín-Gómez A, Marco S, Pardo A. Signal preprocessing in instrument-based electronic noses leads to parsimonious predictive models: application to olive oil quality control. Sensors, 2025. https://doi.org/10.3390/s25030737